You must be signed in to read the rest of this article.
Registration on CDEWorld is free. You may also login to CDEWorld with your DentalAegis.com account.
This article consists of four sections, each of which can be considered a learning objective. The first section defines biofilms and contextualizes them in microbial ecology. The second describes model systems used to investigate biofilms in the laboratory and field. The third part concentrates on specific studies that have significantly contributed to the understanding of biofilm biology. The final section attempts to look into the future and suggests where research might be focused for maximal practical benefit.
What Are Biofilms?
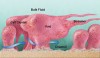
Investigation of biofilms began with the inception of microbial ecology. The Winogradsky column, a clear tube filled with soil in which a visually identifiable stratified microbial community develops, was originally described in the late 19th century and is still a standard learning tool in microbial ecology.1 Although microbial ecologists have worked with biofilms for decades, adoption of the term biofilm has been recent.2 Various definitions occur in the literature, but certain basic concepts overlap. One concept is adherence—the microorganisms form a film on a substratum. Another is the presence of an extracellular matrix that surrounds and encloses the organisms, referred to as extracellular polymeric substance (EPS). This material is comprised of polysaccharides, nucleic acids, and proteins/glycoproteins. An EPS includes not only specific products targeted to the microbial cell surface but also other components exported from living cells. In addition, dead biomass contributes to an EPS though degradation. Laboratory-grown monospecies biofilms (biofilms consisting of a single organism) typically demonstrate topographic variation on a microscopic scale. The biomass can be arranged into upward-developing regions that have been referred to as mushrooms, chimneys, cell clusters, and microcolonies. This arrangement can result in the occurrence of channels or valleys between the taller regions of the biomass. The channels increase the ratio of surface area to biovolume and thereby create a shorter diffusion distance between the nutrient-containing bulk fluid and deeper regions of the biomass. Within the cell clusters, areas devoid of cellular material and presumably filled with EPS can occur. These are known as voids. The fluid overlaying the biofilm, or “bulk fluid,” is distinguished from fluid present within the cell clusters. Under a flow, the biofilm can deform into elongated strands, called streamers, held together by an EPS. Figure 1 provides a diagrammatic overview of biofilm architecture.
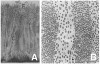
In nature, monospecies biofilms seem to be restricted to certain device-related infections, such as artificial joints, in which a contaminant organism is present prior to introduction or is introduced during the surgery. Otherwise, while a single organism may dominate a biofilm, other organisms always appear. Typically, many organisms coexist in the biofilm community. Classic examples are microbial mats, epilithic biofilms (slime) on submerged rocks, and, the focus of this article, dental plaque. For plaque and other oral biofilms, the adherence sites are enamel, dentine, epithelial surfaces, and reconstructions/appliances. Major components of a supragingival plaque EPS are polymers of glucose and fructose that result from the action of glucosyltransferases produced by streptococci. Smooth-surface supragingival biofilms (those that develop on flat areas of the tooth surface) appear laminar and, under effective oral hygiene practices, are typically no more than a few cell layers thick. They are dominated by streptococci but also contain significant numbers of Veillonella species and Actinomyces species. Small numbers of bacteria that dominate in subgingival communities, especially Porphyromonas species, are also detectable in supragingival biofilms. After a professional cleaning, about 4 hours must pass before cells can be easily found using a microscope; single-cell and two- to three-cell (small) colonies are typical. Even in this early stage, the multicelled colonies can contain more than one type of organism (Figure 2).3 At the gingival margin and in fissures, thicker biofilms can be found. These biofilms display a greater degree of heterogeneity in cellular morphology and organization. A feature known as a palisade (an upward-growing column of cells that is distinct from cells on either side) can be seen in light micrographs or electron micrographs. In some palisade arrangements, the abutting columns are distinguished by cellular morphology, whereas in other arrangements, all columns have cells of similar morphology but with differing degrees of space between the cells within an individual column. Although palisades may be uniform with respect to cell morphology, it is not necessarily true that organisms within the palisade are identical (Figure 3).4
The basis of structure in natural biofilms is the microbial community, ie, the interactions between organisms. In photosynthetic microbial mats, such as those found in hot springs worldwide, vertical stratification is visible to the naked eye. The composition and location of the layers, each of which may contain several major organisms, result from gradients of photosynthetically produced oxygen (Figure 4) and sulfide produced by aerobic sulphate-reducing bacteria.5 The metabolism of the various bacteria proceeds according to a “division of labor,” with one layer passing metabolites to its neighbors. The entire mat can be viewed as a physiologic unit that is the sum of its layers. In supragingival plaque, the relationship between community composition and location within the biofilm is not apparent to the naked eye because biomass is very low and the organisms lack pigmentation. Also, the metabolic relationships between the community subunits are not clearly defined. In the case of thicker supragingival plaque, palisades may be indicative of metabolic interactions. Regardless of the gross morphologic organization of oral biofilms, physiologic interaction is key.
Experimental Biofilms
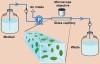
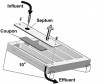
Many model systems exist for experimental investigation of biofilms. Most experimentation is conducted in vitro. Biofilm model systems can be roughly divided according to the manner in which the biofilm contacts the growth medium: by flow of the medium across the biomass or simple diffusion in a static system. A widely used flowing model system is the flowcell (Figure 5A). A bacterial suspension is introduced into a closed channel, one side of which is transparent—normally a microscopy cover slip. After a stagnant attachment period, the medium is continuously flowed through the channel. Adherent biofilm cells are provided with continuous delivery of the fresh medium, and a simultaneous removal of biologic waste and unattached cells occurs. Development of the biofilm adherent to the luminal side of the cover slip is followed using a microscope. The size of flowcells varies considerably. However, as microscopy is used for most data collection, the cells are normally small enough to fit on a standard microscope stage and have two or three channels, roughly 2 mm x 1 mm in cross-sectional area, that extend nearly the entire length of the unit. A similar approach uses small glass capillaries (Figure 5B). Most other flowing-medium approaches involve a device that passes the medium across removable coupons. An example is the drip-flow reactor in which the medium is introduced dropwise onto the elevated end of a coupon that is about the size of a microscope slide and enclosed in a slanted chamber (Figure 5C). The medium flows across the coupon surface by gravity; the biofilm-covered coupon is removed for microscopic analysis and/or destructive analysis of the attached biomass. In this system, a headspace exists over the coupon from which gas samples can be drawn. Coupons can be made of (or coated with) a material appropriate for the biologic system being modeled, eg, stainless steel coupons for investigation of biofilms from food processing plants. Another example includes stirred-vessel reactors, which are typically run as chemostats. Many variations of this type of reactor system exist; the simplest is that in which coupons are simply held suspended on a rack in the stirred medium (Figure 5D). Some designs have circular coupons mounted in a rotating plate at the bottom of the vessel. In the specialized constant-depth film fermenter, the face of the rotating plate is in contact with a sweeper arm and continuously scraped as the plate turns. A biofilm thickness on the coupons is maintained constantly by recessing the coupons into the plate. At each rotation, the sweeper arm shears off the upper biofilm surface. A plug of biofilm develops on the coupon in the recess between the coupon surface and the swept face of the plate. A final example of the flowing model is that of the small cylinders of packed cellulose fibers (similar to cigarette filters), known as Sorbarods. Originally used for orchid culture, they have a very high surface area (the fibers) compressed into a small volume. The filters can be inserted into a tube, such as the barrel of a syringe, and the medium can be pumped through them by applying a head pressure. Flow rates are typically low because of the system back pressure; however, extremely high adherent microbial cell densities (ca 109 cells mL-1) can be attained within the filters. All these flowing models have been applied to oral biofilm research; however, flowcells and rotating disc reactors have been used most commonly.6,7
Static biofilm model systems are typically designed around multiwell plates, 96-well microtiter plates. A bacterial culture is heavily diluted into the fresh medium in the well. After a growth period, the liquid in the wells is removed, leaving behind attached cells on the bottom and sides of the well. Using a plate reader, two types of measurements are often performed for each experiment. A turbidometric reading is taken prior to removal of the well contents, which is a gross measurement of the culture biomass that includes suspended-cell biomass. Then, the liquid is removed, the remaining adherent biofilm stained, and a spectrophotometric reading taken to quantify biofilm biomass in the absence of suspended cells. This latter step is often performed by dissolving the biofilm-bound stain in an appropriate solvent followed by transfer of the eluted stain to a new plate prior to the spectrophotometric measurement. Such a procedure eliminates artifacts associated with uneven biofilm growth and corrects for the inability of the spectrophotometer to detect stained biomass present on the sides of the well. A useful modification is the placement of a coupon within the well; the biofilm thereby can be retrieved intact from the well. For oral biofilms, a disk of hydroxyapatite or of bovine enamel is often used as a removable substratum.8 A variation of the multiwell plate approach is one in which the plate has a removable cover that bears downward-pointing indentations (pins) in the cover surface. When the cover is in place, each pin is immersed in the contents of a well (Figure 5E). The biofilms that develop on the surface of the pins can be easily transferred to a new plate containing a fresh medium, a medium of different composition, or a staining solution—this model is referred to as the Calgary Biofilm Device9 or the Transferable Solid Phase (TSP) model.10
Multiwell plates are also available with a microscopy cover glass as the bottom of the plate; cells attached to the bottom of the well (ie, on the upper surface of the cover glass) can be monitored with an inverted microscope. The typical growth medium in static systems is a 10-fold to 100-fold dilution of the bacterium’s standard growth medium; the lower nutrient concentration seems to promote attachment and reduce growth in the bulk liquid. Saliva sometimes has been used as the diluent,8 and some investigators have used dilute saliva alone as the growth medium.10 In many experiments, the liquid content of the wells is periodically replaced with a fresh medium to support longer growth periods.
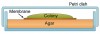
A final example of a stationary system is the colony biofilm (Figure 5F). Here, a membrane filter on an agar-filled Petri dish is inoculated with a bacterial suspension. The growing biofilm is fed by nutrients that diffuse upward through the membrane, and the membrane can be transferred to agars of different composition, including those containing antibiotics. This model is particularly useful for examination of thicker biofilms via cryosectioning followed by microscopy.11 Nearly all of the above-described models have been used to examine biofilms created in vitro with oral bacterial isolates; however, the multiwell plate and the flowcell have been employed most often.
Field (in situ) studies on biofilms in nature have been conducted using methods adopted or, in certain cases, developed by oral microbiologists. Two methodologies of high relevance to oral biofilms are microelectrodes and retrievable substrata. As previously shown in Figure 4, it is possible to follow the development of physiochemical gradients of sulfide and oxygen during light-driven interplay between photosynthetic organisms and sulphate-reducing organisms within the mat.5,12 In oral applications, the use of small electrodes has permitted descriptions of the characteristic short-term drop and subsequent recovery of plaque pH in response to sugar availability—the Stephan curve (Figure 6).13,14 The placement of a substratum in a stream, lake, or soil and its subsequent retrieval has been routinely used by microbial ecologists to obtain biofilm samples from nature15 and undisturbed samples of oral biofilms likewise have been obtained. Typically, a substratum, such as human or bovine enamel, is held in a stent positioned on the mandible or against the hard palate, and chips of enamel also have been enclosed in an orthodontic bracket or some other holder glued to the tooth.16 Various thin pliable films also have been used and are especially good for periodontal applications17 in which stiff bulky substrata are unsatisfactory.
What Has Been Learned from Models?
It is claimed that biofilm cells are intrinsically different in their metabolism and behavior compared with planktonic cells. Historically, the lynchpin of this theory has been antimicrobial/antibiotic resistance: 10-fold to 100-fold greater antibiotic concentrations are required to kill biofilm cells than to kill their planktonic counterparts. Several different mechanisms have been proposed to account for this observation,18 including reduced metabolic activity in biofilm cells. The development of microarrays for gene expression studies has allowed a broad-based search for genes that are regulated by attachment and biofilm growth through a comparison of the planktonic cell transcriptome (the global messenger ribonucleic acid [mRNA] transcript pool) to that of biofilm cells.19 This approach has been employed with Porphyromonas gingivalis20 to show that genes involved in growth processes were down-regulated in biofilms, thereby supporting the general observations of slow metabolic rates in biofilms. Likewise, the transcriptome of Streptococcus mutans was shown to change dependent on biofilm thickness.21 Despite the large number of genes for which changes in expression can be assayed using microarrays, both these studies found fewer than 50 genes were affected. Currently, microarray studies are primarily used as surveys that provide lists of target genes–the true value of microarray analysis will be in the use of global gene expression data to develop testable hypotheses in regard to metabolic behavior of biofilm cells.
In the past decade, cell-to-cell signaling has become recognized as critical to gene transfer and acid adaptation in S mutans. Comparisons of parent and mutant strains have shown an 18-amino acid peptide was critical in biofilm formation and the peptide also possessed bacteriocidal properties.22 This peptide is in a class of molecules known as competence-stimulating peptides (CSPs) that are excreted into culture supernatants. These molecules were identified by their ability to increase deoxyribonucleic acid (DNA) uptake (gene transfer, a process known as competence) when a threshold concentration of the “signal” peptide was reached. CSPs are also known to be involved in the adaptive acid-tolerance response (ATR), in which cells exposed first to a significant but survivable acid shock can subsequently survive an even greater (normally fatal) drop in pH.23 Such molecules seem to encourage biofilm formation: the cell-to-cell transfer of genetic material could be enhanced if distances between cells were minimized. Biofilm cells would benefit from antimicrobial activity directed against other species in nearby locations on the tooth surface, as well as from the ability to tolerate very low pH within dense cell populations of acid-producing organisms.
Studies of biofilms do not need to center on genetics and physiology. The oral care industry has come to realize the benefit of model oral biofilm for testing oral care product formulations and appliances. In these simple comparative studies, laboratory-grown biofilms are used to assess the retention or killing of cells. Recently, chitosan treatment of saliva-coated (intended to mimic the acquired enamel pellicle) substrata has been shown to reduce the attachment rate of certain oral bacteria, as well as to have bacteriostatic properties (causing a reduction in growth rather than outright killing).24 Powered toothbrushes have been evaluated for removal of laboratory-grown biofilms.25 Coupling an ultrasound waveguide to the head of a power toothbrush for mechanical cleaning has been shown to be more effective at biofilm removal than either the power toothbrush alone or the application of ultrasound alone.26 A mixed-species in-vitro oral biofilm model has been used to show the relationship between antimicrobial penetration into the biofilm, antimicrobial mode of action, and bacterial membrane integrity.27
Most investigations of oral biofilm biology have relied on pure cultures of organisms. However, it is accepted that the oral cavity is home to at least 700 different bacteria.28 Because oral biofilms are inherently polymicrobial, experiments involving multiple bacteria likely better represent the ecology of oral biofilms than do experiments based on a single strain. Likewise, the use of medium formulations that approach the composition of saliva or gingival crevicular fluid likely better mimic conditions within the oral cavity than does the use of the complex media typical in the culture for single bacterial strains. Experiments in a flowcell system using saliva as the sole nutrient source have indicated that Streptococcus gordonii strain DL1 can grow as a monoculture biofilm, whereas Streptococcus oralis strain 34 and Actinomyces oris (formerly Actinomyces naeslundii) strain T14V cannot. However, when grown together, S oralis 34 and A oris T14V cooperate to produce large amounts of biofilm biomass (Figure 7). In contrast, the combination of S gordonii DL1 with either S oralis 34 or A oris T14V has no synergistic effect: S gordonii grows just as it does in monoculture whereas neither of the other bacteria grows.6 Further work suggests the mutualistic interaction between S oralis and A oris is, in part, a result of autoinducer 2 (AI-2) signaling between the cells. Like CSPs, autoinducers are low-molecular-weight molecules that are excreted into the medium and effect gene expression, and thus metabolism, in organisms that can sense the autoinducer.29 If a mutant of S oralis 34 unable to synthesize AI-2 is grown as a saliva-fed biofilm together with A oris T14V, the mutualistic growth fails to occur.30 When AI-2 synthesis in the mutant S oralis is restored by insertion of a plasmid bearing the AI-2 synthesis gene, mutualistic growth is restored. The addition of AI-2 to the saliva likewise restores mutualistic growth. These data clearly indicate a role for AI-2 signaling in mutualistic growth; however, a mechanism (the downstream consequences of the signaling process) has yet to be revealed. Flowcell-based models of interorganismal interactions in oral biofilms are not limited to bacteria. Research has recently shown AI-2 signaling by S gordonii is a positive regulator of biofilm formation by Candida albicans31—an intriguing observation whereby a signal of bacterial origin modulates physiology in a higher organism.
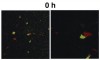
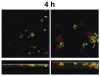
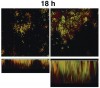
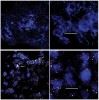
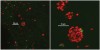
Microscopic observations have defined the spatiotemporal relationship between certain microorganisms in vivo, and physiologic interactions exist that could account for the spatial relationships. For example, veillonellae cannot use sugars for growth—they require short-chain organic acids, such as lactic acid. Streptococci use sugars from which they produce organic acids during growth. Thus, in vitro investigations of physiologic interaction between these bacteria have been popular. A gnotobiotic rat model of oral bacterial colonization and growth used more than two decades ago32,33 yielded in vivo data supportive of interaction between these bacteria, yet data on this interaction in humans have been acquired only recently. Veillonellae were found in immediate proximity to streptococci in early supragingival plaque, and composition of the veillonellae community was shown to change dramatically in 4 hours (Figure 8).34 Spatial relationships between organisms also have been noted at a gross scale in the periodontal pocket.35 In this study, teeth and associated tissue were extracted from patients with advanced periodontitis. Immunohistochemistry performed on tissue serial sections that included the tooth surface as well as the associated sulcus revealed that Actinomyces naeslundii was primarily at the tooth surface in the upper regions of the pocket. Campylobacter rectus was present as large clumps in the deepest regions of the pocket and, therefore in close proximity to the tooth surface and the epithelium. P gingivalis, in contrast, was distributed as small aggregates throughout the periodontal pocket and could be seen in close proximity to the epithelial surface. In a similar approach using samples from patients with refractory periapical periodontitis,36 P gingivalis was seen scattered throughout the periradicular biofilm. Fusobacterium nucleatum was found in the middle of the biofilm (between the tooth surface and superficial surface of the biofilm), and Tannerella forsythensis was present primarily in the superficial layer. The positioning of organisms within the periodontal pocket and in other interfacial areas between host tissue and tooth likely reflects gradients of oxygen, gingival crevicular fluid, and saliva. However, the broad spatial distribution of P gingivalis in these situations suggests that this organism is better adapted to grow throughout a range of environmental conditions.
How are these spatial relationships established? Environmental conditions are clearly an important driver; however, other factors contribute. Forty years ago, researchers discovered that oral bacterial isolates bind to oral bacteria of other genera or species37—a phenomenon that is known as coaggregation. This characteristic, which is widespread among oral bacteria, is assayed in vitro by mixing suspensions of the two cultured candidate organisms. If they are coaggregation partners, floculation (formation of coaggregates) occurs. By adding a third bacterium that recognizes the other two, it is possible to create coaggregates composed of bacterial strains that do not coaggregate on their own.38 This complex framework has led to the development of a model that approximates the bacterial succession that is known to occur in oral biofilms.39 Thus, the hypothesis that coaggregation is important in plaque formation is attractive. However, coaggregation is an in vitro phenomenon (more properly, an assay)—the significance of which in vivo is difficult to assess. The phylogenetic identification of a tooth surface-bound organism in situ, even to the species level, does not describe its coaggregation profile; instead, the profile is determined by the presence of adhesins or receptors on the cell surface. The presence of these molecules has, in the best circumstances, a complex congruence with bacterial taxonomy. Little is known about the adhesins; however, many coaggregation interactions, especially those between the initial colonizers in supragingival biofilms, are mediated by a cell-surface polysaccharide called receptor polysaccharide (RPS), borne on certain streptococci. This molecule participates in a lectinlike recognition with yet-to-be characterized protein adhesins on partner organisms. In one case, enough is known about the adhesin for a coaggregation-mediated interaction to be sought in situ: the interaction of RPS-bearing streptococci with Actinomyces species that bear type 2 fimbriae. Antibodies specific to RPS and type 2 fimbriae were used in a retrievable enamel chip model to show the immediate proximity of cells bearing these antigens (Figure 9),3 thereby providing strong evidence that an adhesin–receptor interaction could have created this spatial relationship within natural human plaque. However, definitive evidence would require those very organisms (the antibody-reactive cells that are juxtaposed) to be isolated and shown to be coaggregation partners using the in vitro assay. This proof was obtained, but in a somewhat different manner: an RPS-bearing streptococcal cell was micromanipulated from the enamel surface and an antibody-unreactive streptococcal cell was co-isolated, ie, the two were in immediate proximity. After culturing, it could be shown that the unreactive cell did, in fact, coaggregate with the RPS-bearing streptococcus.10
What Does the Future Hold?
In the past 15 years, biofilm research has undergone explosive growth as a specialty area within microbiology. A recent international congress on biofilm biology attracted approximately 600 attendees from all microbiologic backgrounds.40 When biofilm systems from such diverse environments as plants, rocks, and teeth are examined from a unified perspective, then understanding of these systems is synergistically advanced. Areas of concentration in the next decade will include the extracellular matrix, gene expression in multispecies biofilms, and, particularly for oral micro-organisms, the consequences of coaggregation. Experimentation will shift towards complex models that incorporate eukaryotic cells (as in oral bacterial invasion model systems),41 examine host response to biofilms from a genomic standpoint,42 and are based on nutrient compositions similar to those in situ. Furthermore, biofilm growth and metabolism under anaerobic conditions are highly relevant to many environmental and medical situations—this area also should become prominent. As a field of study, biofilm microbiology is young enough that hypotheses can be simple, while being old enough that a sufficient body of evidence exists on which detailed hypotheses can be built. Periodontal disease has long been investigated from a microbial ecological perspective,43 and it is now time to augment those data from a biofilm perspective. Caries has been postulated to arise from shifts within microbial communities in a type of feedback loop involving ecologic parameters;44 biofilm models now can be used to test these hypotheses in vitro and in vivo and be employed to assess treatment and prevention strategies. Other oral diseases such as candidiasis are beginning to be explored with biofilm biology as the scaffold. Finally, disease does not need to be the focus. Prior to disease, health reigns. What exactly is a healthy oral microflora? Can the microflora characteristic of oral diseases be returned to the healthy state without ecologically detrimental treatments, such as antibiotics? Can a healthy microflora be stimulated to prevent disease? These questions revolve around interactions between the bacterium and environment, as well as interactions between the bacteria themselves. Oral biofilm microbiologists have a solid foundation on which they can build, a foundation that is envied by biofilm researchers in many other investigative areas. Oral biofilms are paradigms.
References
1. Institute of Cell and Molecular Biology at The University of Edinburgh Web site. The microbial world. Winogradsky column: perpetual life in a tube. http://www.biology.ed.ac.uk/research/groups/jdeacon/microbes/winograd.htm.
Accessed October 19, 2009.
2. Palmer RJ Jr. Peter Hirsch and biofilms: microbial ecology’s role in a “new” field. Microb Ecol. 2004;47(3):200-204.
3. Palmer RJ Jr, Gordon SM, Cisar JO, et al. Coaggregation-mediated interactions of streptococci and actinomyces detected in initial human dental plaque. J Bacteriol. 2003;185(11):3400-3409.
4. Listgarten MA. The structure of dental plaque. Periodontol 2000. 1994;5:552-565.
5. Jørgensen BB, Revsbech NP, Cohen Y. Photosynthesis and structure of benthic microbial mats: microelectrode and SEM studies of four cyanobacterial communities. Limnol Oceanogr. 1983;28(6):1075-1093.
6. Palmer RJ Jr, Kazmerzak K, Hansen MC, et al. Mutualism versus independence: strategies of mixed-species oral biofilms in vitro using saliva as the sole nutrient source. J Bacteriol. 2001;69(9):5794-5804.
7. Pratten J. Growing oral biofilms in a constant depth film fermentor (CDFF). Curr Protoc Microbiol. 2007 Aug;Chapter 1:Unit 1B.5.
8. Guggenheim M, Shapiro S, Gmür R, et al. Spatial arrangements and associative behavior of species in an in vitro oral biofilm model. Appl Environ Microbiol. 2001;67(3):1343-1350.
9. Harrison J, Ceri H, Yerly J, et al. The use of microscopy and three-dimensional visualization to evaluate the structure of microbial biofilms cultivated in the Calgary Biofilm Device. Biol Proced Online. 2006;8(1):194-215.
10. Chalmers NI, Palmer RJ Jr, Cisar JO, et al. Characterization of a Streptococcus sp.-Veillonella sp. community micromanipulated from dental plaque. J Bacteriol. 2008;190(24):8145-8154.
11. Rani SA, Pitts B, Beyenal H, et al. Spatial patterns of DNA replication, protein synthesis, and oxygen concentration within bacterial biofilms reveal diverse physiological states. J Bacteriol. 2007;189(11):4223-4233.
12. Jørgensen BB, Revsbech NP, Blackburn TH, et al. Diurnal cycle of oxygen and sulfide microgradients and microbial photosynthesis in a cyanobacterial mat sediment. Appl Environ Microbiol. 1979;38(1):46-58.
13. Fejerskov O, Scheie AA, Manji, F. The effect of sucrose on plaque pH in the primary and permanent dentition of caries-inactive and -active Kenyan children. J Dent Res. 1992; 71(1):25-31.
14. Stephan RM. Changes in hydrogen-ion concentration on tooth surfaces and in carious lesions. J Am Dent Assoc. 1940;27:718-723.
15. Hirsch P, Pankratz SH. Study of bacterial populations in natural environments by use of submerged electron microscope grids. Z Allg Mikrobiol. 1970;10(8):589-605.
16. Manning RH, Edgar WM. Intra-oral models for studying de- and remineralization in man: methodology and measurement. J Dent Res. 1992;71(spec no):895-900.
17. Wecke J, Kersten T, Madela K, et al. A novel technique for monitoring the development of bacterial biofilms in human periodontal pockets. FEMS Microbiol Lett. 2000;191(1):95-101.
18. Drenkard E. Antimicrobial resistance of Pseudomonas aeruginosa biofilms. Microbes Infect. 2003;5(13):1213-1219.
19. Goodman AL, Lory S. Analysis of regulatory networks in Pseudomonas aeruginosa by genomewide transcriptional profiling. Curr Opin Microbiol. 2004;7(1):39-44.
20. Lo A, Seers CA, Boyce, JD, et al. Comparative transcriptomic analysis of Porphyromonas gingivalis biofilm and planktonic cells. BMC Microbiol. 2009;9(1):18.
21. Shemesh M, Tam A, Kott-Gutkowski M, et al. DNA-microarrays identification of Streptococcus mutans genes associated with biofilm thickness. BMC Microbiol. 2008;8(1):236.
22. Petersen FC, Fimland G, Scheie AA. Purification and functional studies of a potent modified quorum-sensing peptide and a two-peptide bacteriocin in Streptococcus mutans. Mol Microbiol. 2006;61(5):1322-1334.
23. Li YH, Hanna MN, Svensäter G, et al. Cell density modulates acid adaptation in Streptococcus mutans: implications for survival in biofilms. J Bacteriol. 2001; 183(23):6875-6884.
24. Busscher HJ, Engels E, Dijkstra RJ, et al. Influence of a chitosan on oral bacterial adhesion and growth in vitro. Eur J Oral Sci. 2008;116(5):493-495.
25. Adams H, Winston MT, Heersink J, et al. Development of a laboratory model to assess the removal of biofilm from interproximal spaces by powered tooth brushing. Am J Dent. 2002;15 spec no:12B-17B.
26. Mourad PD, Roberts FA, McInnes C. Synergistic use of ultrasound and sonic motion for removal of dental plaque bacteria. Compend Contin Ed Dent. 2007;28(7):354-358.
27. Takenaka S, Trivedi HM, Corbin A, et al. Direct visualization of spatial and temporal patterns of antimicrobial action within model oral biofilms. Appl Environ Microbiology. 2008;74(6):1869-1875.
28. Aas JA, Paster BJ, Stokes LN, et al. Defining the normal bacterial flora of the oral cavity. J Clin Microbiol. 2005;43(11):5721-5732.
29. Bassler BL. How bacteria talk to each other: regulation of gene expression by quorum sensing. Curr Opin Microbiol. 1999;2(6):582-587.
30. Rickard AH, Palmer RJ Jr, Blehert DS, et al. Autoinducer 2: a concentration-dependent signal for mutualistic bacterial biofilm growth. Mol Microbiol. 2006;60(6):1446-1456.
31. Bamford CV, d'Mello A, Nobbs AH, et al. Streptococcus gordonii modulates Candida albicans biofilm formation through intergeneric communication. Infect Immun. 2009;77(9):3696-3704.
32. Mikx FH, van der Hoeven JS, König KG, et al. Establishment of defined microbial ecosystems in germ-free rats. I. The effect of the interactions of Streptococcus mutans or Streptococcus sanguis with Veillonella alcalescens on plaque formation and caries activity. Caries Res. 1972;6(3):211-223.
33. van der Hoeven JS, Toorop AI, Mikx RH. Symbiotic relationship of Veillonella alcalescens and Streptococcus mutans in dental plaque in gnotobiotic rats. Caries Res. 1978;12(3):142-147.
34. Palmer RJ Jr, Diaz PI, Kolenbrander PE. Rapid succession within the Veillonella population of a developing human oral biofilm in situ. J Bacteriol. 2006;188(11):4117-4124.
35. Noiri Y, Li L, Ebisu S. The localization of periodontal-disease-associated bacteria in human periodontal pockets. J Dent Res. 2001;80(10): 1930-1934.
36. Noguchi N, Noiri Y, Narimatsu M, et al. Identification and localization of extraradicular biofilm-forming bacteria associated with refractory endodontic pathogens. Appl Environ Microbiol. 2005;71(12):8738-8743.
37. Gibbons RJ, Nygaard M. Interbacterial aggregation of plaque bacteria. Arch Oral Biol. 1970;15(12):1397-1400.
38. Kolenbrander PE, Andersen RN, Holdeman LV. Coaggregation of oral Bacteroides species with other bacteria: central role in coaggregation bridges and competitions. Infect Immun. 1985;48(3):741-746.
39. Kolenbrander PE, London J. Adhere today, here tomorrow: oral bacterial adherence. J Bacteriol. 1993;175(11):3247-3252.
40. Palmer RJ Jr, Stoodley P. Biofilms 2007: broadened horizons and new emphases. J Bacteriol. 2007;189(22):7948-7960.
41. Yılmaz O, Verbeke P, Lamont RJ, et al. Intercellular spreading of Porphyromonas gingivalis infection in primary gingival epithelial cells. Infect Immun. 2006;74(1):703-710.
42. Handfield M, Baker HV, Lamont RJ. Beyond good and evil in the oral cavity: insights into host-microbe relationships derived from transcriptional profiling of gingival cells. J Dent Res. 2008;87(3):203-223.
43. Socransky SS, Haffajee AD. Periodontal microbial ecology. Periodontol 2000. 2005;38:135-187.
44. Marsh PD. Are dental diseases examples of ecological catastrophes? Microbiology. 2003;149(2):279-294.
About the Author
Robert J. Palmer Jr, PhD, Staff Scientist–Microbiologist, Oral Infection and Immunity Branch, National Institute of Dental Craniofacial Research at the National Institutes of Health, Bethesda, Maryland